Mar 18, 2026
Model Medicines’ AmesNet™Achieves Best-in-Class Performance in AI-Driven Ames Mutagenicity Prediction
Novel Task-Conditioned Learning architecture outperforms FDA, MIT, Baidu, and University of Sydney models in both sensitivity and balanced accuracy on out-of-domain chemical data; extends GALILEO™ platform from drug discovery into ADMET safety prediction

SAN DIEGO, CA — March 11, 2026 — Model Medicines today announced that AmesNet™, a deep learning model for predicting Ames test mutagenicity, demonstrated best-in-class performance across all benchmarked approaches in predicting chemical safety on out-of-domain data. AmesNet outperformed models from major organizations, including the FDA's DeepAmes (FDA/NCTR), ChemProp (MIT), GROVER (Baidu Research), and the University of Sydney. The model uses a new AI architecture called Task-Conditioned Learning (TCL) and is built on the Company's advanced AI platform, GALILEO™.
Key Highlights:
Industry Problem: The Ames test is a required genotoxicity assay for all novel small-molecule therapeutics prior to human clinical trials. Regulatory agencies have moved to enable AI alternatives. The Second Ames/QSAR International Challenge Project (2020–2022, 21 teams, 11 countries) yielded an average participant sensitivity of only 0.46 on out-of-domain data.
Best-in-Class Performance: AmesNetTM achieves a sensitivity of 0.73 and a balanced accuracy of 0.81 on out-of-domain test data (n = 4,528). A sensitivity improvement up to 46% over existing approaches.
New Modeling Paradigm: Task-Conditioned Learning (TCL) architecture captures assay-context-dependent mutagenicity that all prior “Unconditioned” models either don’t account for or average away.
Early-Stage Screening Enabled: AmesNet’s performance enables high-confidence Ames screening at scale during early discovery, pulling preclinical safety assessment forward from a late-stage bottleneck to a continuous input in compound prioritization.
Platform Extension: AmesNetTM extends GALILEO™ from target-based drug discovery to ADMET safety prediction.
Open Science: The model, code, and data are publicly available at github.com/Model-Medicines/TCL-Ames
Preprint: the preprint for AmesNetTM can be found on bioRxiv - https://www.biorxiv.org/content/10.1101/2025.03.20.644379v2
“The Ames test is one of the most consequential checkpoints in drug development. It has remained a late-stage gate because no computational model has been reliable enough to deploy at program concept,” said Daniel Haders, PhD, CEO of Model Medicines. “AmesNet changes that equation. Task-Conditioned Learning delivers the sensitivity and balanced accuracy needed to screen compound libraries with confidence at the earliest stages of discovery. This is what it looks like when a platform built for drug discovery extends into drug safety.”
Background: The Ames Testing Bottleneck
The Ames test is a required genotoxicity assay for all novel small-molecule therapeutics prior to human clinical trials. GLP-compliant testing often exceeds $10,000 per compound and requires approximately 2 grams of material, making routine screening impractical during early discovery. Developers typically defer testing until regulatory submission, at which point tens of millions of dollars have been sunk into programs. Regulatory agencies have moved to enable AI alternatives: the FDA Modernization Act provides a legal framework for computational models to replace wet-lab testing, the FDA’s AI4TOX/SafetAI initiatives target AI for toxicological endpoints, and ICH M7 endorses computational Ames prediction. However, no model has met this mandate. The Second Ames/QSAR International Challenge Project (2020–2022, 21 teams, 11 countries) yielded an average participant sensitivity of only 0.46 on out-of-domain data. The FDA’s DeepAmes achieved 0.47; when tuned to 0.87 sensitivity, balanced accuracy fell to 0.52; essentially random classification.
AmesNet™ Benchmark Results
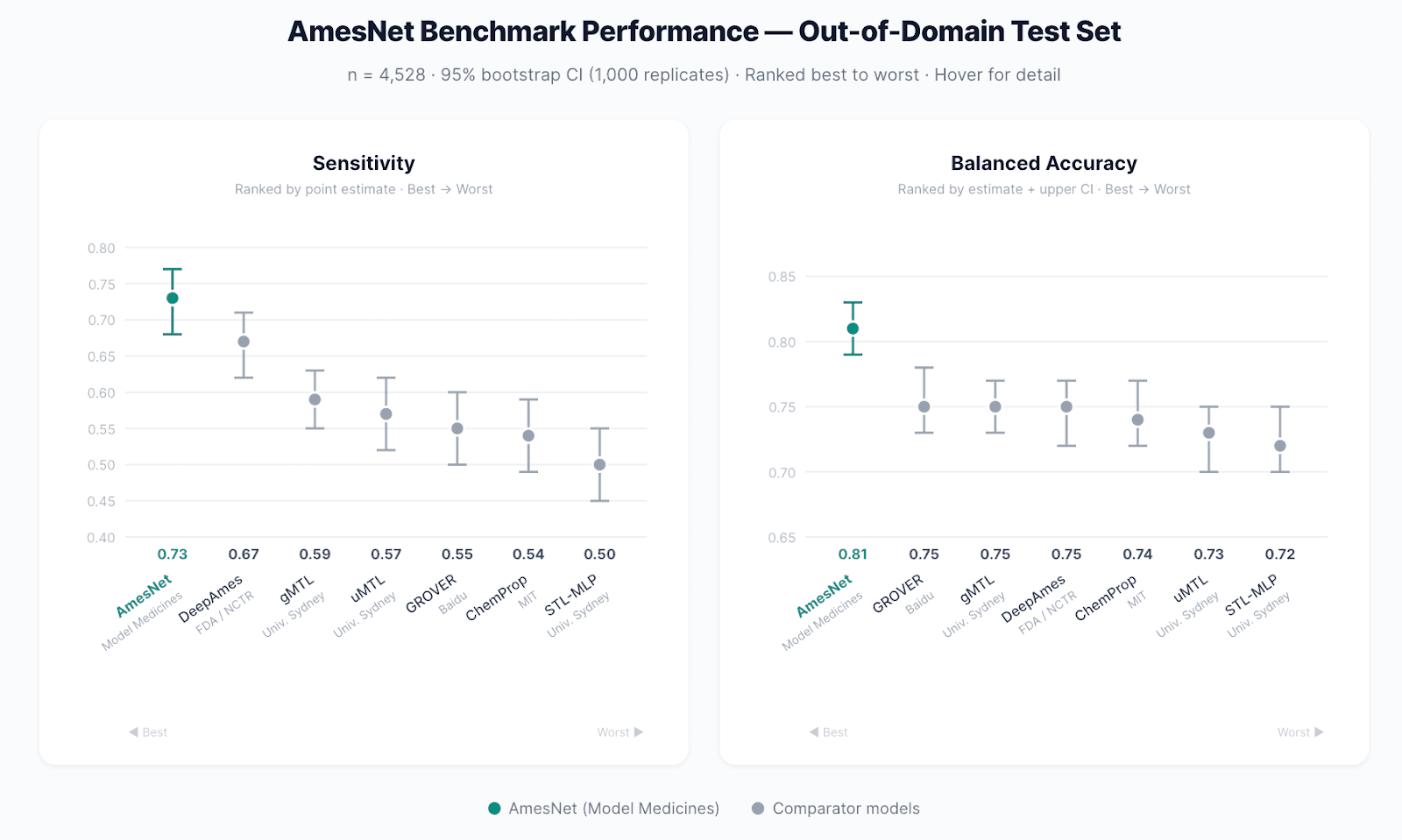
AmesNet™ was evaluated on a withheld out-of-domain test set of 4,528 data points comprising compounds chemically dissimilar from the training data:
Model | Source | Sensitivity (95% CI) | Balanced Accuracy (95% CI) |
AmesNet | Model Medicines | 0.73 (0.68–0.77) | 0.81 (0.79–0.83) |
STL-DeepAmes | FDA / NCTR | 0.67 (0.62–0.71) | 0.75 (0.72–0.77) |
gMTL-MLP | Univ. of Sydney | 0.59 (0.55–0.63) | 0.75 (0.73–0.77) |
uMTL-MLP | Univ. of Sydney | 0.57 (0.52–0.62) | 0.73 (0.70–0.75) |
STL-GROVER | Baidu Research | 0.55 (0.50–0.60) | 0.75 (0.73–0.78) |
STL-ChemProp | MIT | 0.54 (0.49–0.59) | 0.74 (0.72–0.77) |
STL-MLP | Univ. of Sydney | 0.50 (0.45–0.55) | 0.72 (0.70–0.75) |
AmesNet exhibits non-overlapping 95% confidence intervals against nearly all comparators on both metrics. Sensitivity improvement ranges from 9% over DeepAmes to 46% over STL-MLP.
“Every existing AI model for Ames prediction either ignores the structure of the assay itself or handles it through multitask modeling. They produce predictions without conditioning on bacterial strain or metabolic activation,” said Tyler Umansky, Senior Manager, Head of Platform and Machine Learning, and lead author on the paper. “AmesNet conditions on these variables because the biology demands it. The result is a 46% improvement in sensitivity over the baseline and recovery of mutagenic compound classes that every other model systematically misses.”
Task-Conditioned Learning (TCL)
The Ames test is a battery of experiments across multiple bacterial strains. Each strain is sensitive to a different type of DNA mutation, conducted with and without a liver enzyme fraction (S9) that simulates human drug metabolism. A compound may be mutagenic in one strain but not another, or only when metabolized. All prior AI Models are “Unconditioned” and produce a single prediction without conditioning on strain or metabolic activation. AmesNet’s TCL architecture resolves this issue with a dual-branch design: one branch encodes molecular structure; the second encodes assay conditions (strain identity and ±S9 status). The model learns separate decision boundaries for each context rather than averaging across all conditions.
The conditioning channel was injected into two third-party encoders - ChemProp (MIT) and GROVER (Baidu) - to confirm that TCL drives performance gain. TCL-ChemProp improved sensitivity from 0.54 to 0.65 (+20%) with no trade-off in balanced accuracy. Notably, AmesNet still outperformed all TCL encoder-swap variants. This demonstrated that GALILEO™’s molecular representation provides a proprietary compounding advantage on top of the conditioning framework.
Correctly identifying mutagenic compounds, known as sensitivity, is the most critical metric in AI-driven Ames testing. False negatives allow dangerous compounds to advance undetected. Existing models fail on sensitivity because compound classes such as aromatic amines (requiring S9 activation) and planar intercalators (detectable only by frameshift-sensitive strains) produce context-dependent signals that Unconditioned Models dilute. Structural enrichment analysis confirms AmesNetTM recovers these classes: planar aromatic intercalators (n = 27–46), aromatic amines (n = 20–33), and aromatic N-heterocycles (n = 17–25).
A Paradigm Shift for Ames Screening and Preclinical Development.
A model with AmesNet's sensitivity and balanced accuracy in novel chemical space enables high-confidence mutagenicity screening at scale during early drug discovery. This shifts Ames testing from a wet-lab, late-stage gate on a single candidate to a computational filter applied across entire compound libraries at program concept. This repositioning advances preclinical safety assessment by years in the development timeline, and transforms Ames screening from a late-stage binary pass/fail checkpoint into a continuous, scalable input for compound prioritization. AmesNetTM enables developers to eliminate mutagenic liabilities from their pipelines before they become costly.
GALILEO™: One Platform, Discovery Through Development
AmesNet™ is not a standalone product. It is the latest capability within GALILEO™, the AI platform Model Medicines uses to discover, optimize, and de-risk its own wholly owned drug pipeline. The company is progressing two drug candidates towards IND. The first, MDL-001, is a first-in-class, best-in-class oral pan-antiviral with demonstrated efficacy against representative viruses within 6 viral families (Influenza, RSV, Covid, HCV, HBV, HDV). The company is targeting an IND submission in 2H 2026. The second, MDL-4102, targets BRD4, a previously "undruggable" transcriptional control target with robust preclinical activity. The company is targeting an IND submission in 2027.
Additional Preprints:
MDL-001: An Oral, Safe, and Well-Tolerated Broad-Spectrum Inhibitor of Viral Polymerases
ChemPrint: An AI-Driven Framework for Enhanced Drug Discovery
About Model Medicines
Model Medicines is an AI-first biotechnology company engineering first-in-class small molecules that target the biological linchpins underlying disease. The company’s research spans infectious disease, oncology, and inflammation, with programs designed around conserved molecular choke points that drive multiple pathologies. Model Medicines has discovered a direct-acting, non-nucleoside, broad-spectrum antiviral (MDL-001) and a BRD4 inhibitor with no measurable activity against BRD2/3 (MDL-4102). Its work demonstrates how large-scale computation can uncover entirely new classes of drugs once thought unreachable. Model Medicines is advancing a new generation of therapeutics that redefine what is possible in modern drug discovery. Learn more at www.modelmedicines.com
Details
Date
Category
Preprint
Reading
3 Mins
Author

Patrick ONeill
Investor Relations
Related News


